models.logentropy_model – LogEntropy model¶
This module allows simple Bag of Words (BoW) represented corpus to be transformed into log entropy space. It implements Log Entropy Model that produces entropy-weighted logarithmic term frequency representation.
Empirical study by Lee et al. 2015 1 suggests log entropy-weighted model yields better results among other forms of representation.
References
- 1
Lee et al. 2005. An Empirical Evaluation of Models of Text Document Similarity. https://escholarship.org/uc/item/48g155nq
- class gensim.models.logentropy_model.LogEntropyModel(corpus, normalize=True)¶
Bases:
TransformationABCObjects of this class realize the transformation between word-document co-occurrence matrix (int) into a locally/globally weighted matrix (positive floats).
This is done by a log entropy normalization, optionally normalizing the resulting documents to unit length. The following formulas explain how o compute the log entropy weight for term
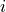 in document
in document 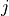 :
: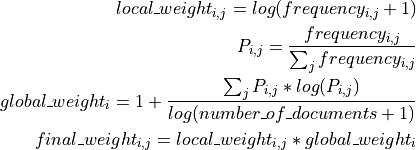
Examples
>>> from gensim.models import LogEntropyModel >>> from gensim.test.utils import common_texts >>> from gensim.corpora import Dictionary >>> >>> dct = Dictionary(common_texts) # fit dictionary >>> corpus = [dct.doc2bow(row) for row in common_texts] # convert to BoW format >>> model = LogEntropyModel(corpus) # fit model >>> vector = model[corpus[1]] # apply model to document
- Parameters
corpus (iterable of iterable of (int, int)) – Input corpus in BoW format.
normalize (bool, optional) – If True, the resulted log entropy weighted vector will be normalized to length of 1, If False - do nothing.
- add_lifecycle_event(event_name, log_level=20, **event)¶
Append an event into the lifecycle_events attribute of this object, and also optionally log the event at log_level.
Events are important moments during the object’s life, such as “model created”, “model saved”, “model loaded”, etc.
The lifecycle_events attribute is persisted across object’s
save()andload()operations. It has no impact on the use of the model, but is useful during debugging and support.Set self.lifecycle_events = None to disable this behaviour. Calls to add_lifecycle_event() will not record events into self.lifecycle_events then.
- Parameters
event_name (str) – Name of the event. Can be any label, e.g. “created”, “stored” etc.
event (dict) –
Key-value mapping to append to self.lifecycle_events. Should be JSON-serializable, so keep it simple. Can be empty.
This method will automatically add the following key-values to event, so you don’t have to specify them:
datetime: the current date & time
gensim: the current Gensim version
python: the current Python version
platform: the current platform
event: the name of this event
log_level (int) – Also log the complete event dict, at the specified log level. Set to False to not log at all.
- initialize(corpus)¶
Calculates the global weighting for all terms in a given corpus and transforms the simple count representation into the log entropy normalized space.
- Parameters
corpus (iterable of iterable of (int, int)) – Corpus is BoW format
- classmethod load(fname, mmap=None)¶
Load an object previously saved using
save()from a file.- Parameters
fname (str) – Path to file that contains needed object.
mmap (str, optional) – Memory-map option. If the object was saved with large arrays stored separately, you can load these arrays via mmap (shared memory) using mmap=’r’. If the file being loaded is compressed (either ‘.gz’ or ‘.bz2’), then `mmap=None must be set.
See also
save()Save object to file.
- Returns
Object loaded from fname.
- Return type
object
- Raises
AttributeError – When called on an object instance instead of class (this is a class method).
- save(fname_or_handle, separately=None, sep_limit=10485760, ignore=frozenset({}), pickle_protocol=4)¶
Save the object to a file.
- Parameters
fname_or_handle (str or file-like) – Path to output file or already opened file-like object. If the object is a file handle, no special array handling will be performed, all attributes will be saved to the same file.
separately (list of str or None, optional) –
If None, automatically detect large numpy/scipy.sparse arrays in the object being stored, and store them into separate files. This prevent memory errors for large objects, and also allows memory-mapping the large arrays for efficient loading and sharing the large arrays in RAM between multiple processes.
If list of str: store these attributes into separate files. The automated size check is not performed in this case.
sep_limit (int, optional) – Don’t store arrays smaller than this separately. In bytes.
ignore (frozenset of str, optional) – Attributes that shouldn’t be stored at all.
pickle_protocol (int, optional) – Protocol number for pickle.
See also
load()Load object from file.
